Classification in Cryo-Electron Tomograms
SHREC 2019 Track

Motivation
There is a noticeable gap in knowledge about the organization of cellular life at the mesoscopic level. With the advent of the direct electron detectors and the associated resolution revolution, cryo-electron tomography (cryo-ET) has the potential to bridge this gap by simultaneously visualizing the cellular architecture and structural details of macromolecular assemblies, thee-dimensionally. The technique offers insights in key cellular processes and opens new possibilities for rational drug design. However, the biological samples are radiation sensitive, which limits the maximal resolution and signal-to-noise ratio. Innovation in computational methods remains key to derive biological information from cryo-electron tomograms.
Task
In this SHREC track, we propose a task of localization (e.g. particle picking) and classification (e.g. template matching) of biological particles in the cryo-electron tomogram volume. We provide physically accurately simulated cryo-electron tomograms and ground truth for all of them except the test tomogram. We hope that this will enable researchers to try out different methods, including machine learning and statistical approaches.
Dataset
To provide participants with as accurate ground truth information as possible, we have created a physically accurate simulator that is able to generate a realistic cryo electron tomogram.
The dataset consists of 10 tomograms, with 1nm/voxel resolution, with a size of 512x512x512 voxels. Each tomogram is packed with on average 2500 bio-particles of 12 uniformly distributed and rotated different proteins, various in size and structure, with the following PDB identificators:
For each tomogram but the test tomogram, we also provide projections from which reconstruction was done, a ground truth volume, from which the tomogram was initially modeled and also a file with locations of particles and their true class.
Download datasets PaperRegistration
To participate in the track, please send us an email. In it, please confirm your interest in participation and if applicable, please also mention your affiliation and co-authors.
Submission
From participants, no later than the deadline mentioned in the schedule, we expect results submitted along with a one-page description of the method used to generate them. Results should be presented as a .txt file containing the found particles, in the similar fashion to the ground truth text files. Data should be formatted in 4 columns: predicted class (PDB ID), estimated center Z coordinates, estimated center Y coordinates, estimated center Z coordinates.
Evaluation
The main goal of the track is to localize and classify biological particles in the tomogram. The performance of methods will be evaluated solely on the test tomogram: the only tomogram for which ground truth is not provided. Following metrics will be measured and compared: precision, recall, F1 score.
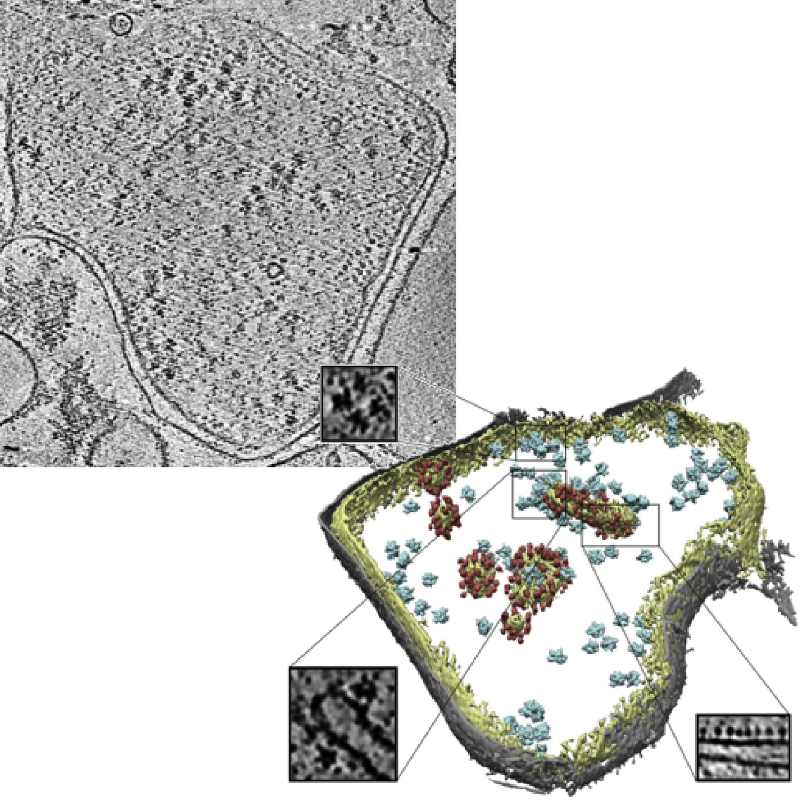
Slice of a tomogram and a 3D visualization obtained from the tomogram.
Image credits: Pfeffer S, Woellhaf MW, Herrmann JM, Förster F, Organization of the mitochondrial translation machinery studied in situ by cryoelectron tomography. Nature communications 6:6019 (2015)
Organizers
- Ilja Gubins 1
- Gijs van der Schot 2
- Remco C. Veltkamp 1
- Friedrich G. Forster 2
2: Utrecht Universty, Department of Chemistry
Contact us
Schedule
The registration and submission deadlines are in AoE (Anywhere on Earth) timezone.
| February |
Registration deadline |
| March |
Submission deadline of the results |
| March 15 | Track submission to SHREC for review |
| March 26 | Reviews are done, notification to the participants |
| April 5 | Camera-ready track paper is submitted for inclusion into the proceedings |
| May 5-6 | Eurographics 2019 + 3DOR + SHREC |